MOLECULAR DETECTION
Conventional isolation methods generally include a cultivation step and molecular methods can therefore be faster for the detection of fungi from spoiled foods. Theoretically, in PCR-based methods, small amounts of the target organism can be detected by amplifying their DNA in a considerably short time frame. Next generation DNA sequencing techniques are becoming more common, but most assays rely on conventional and real-time PCR methods. A compact overview of these methods is given below.
DNA EXTRACTION
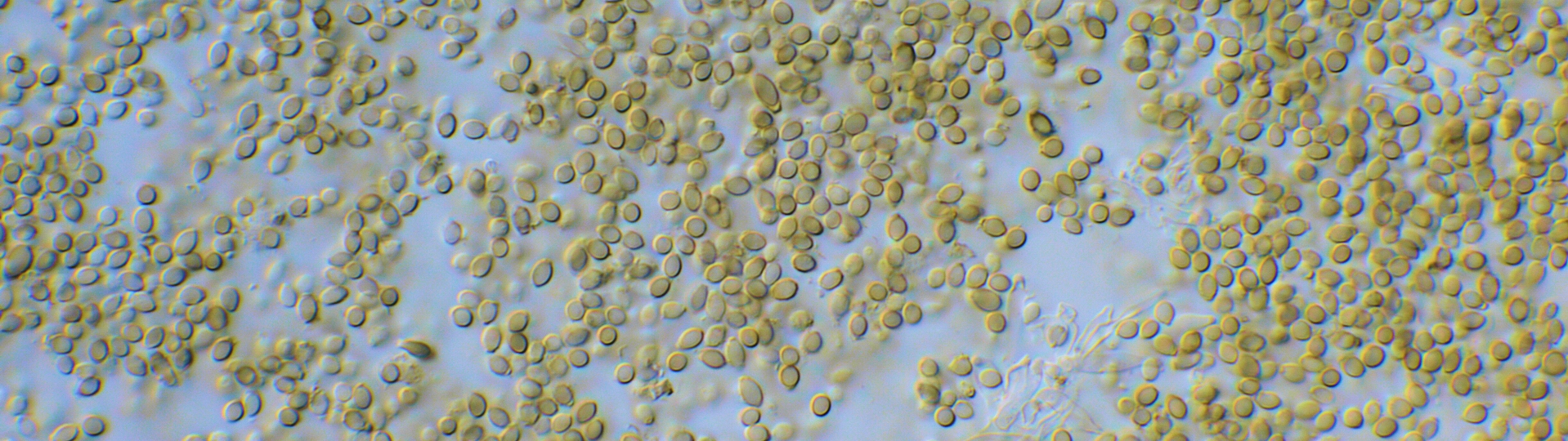
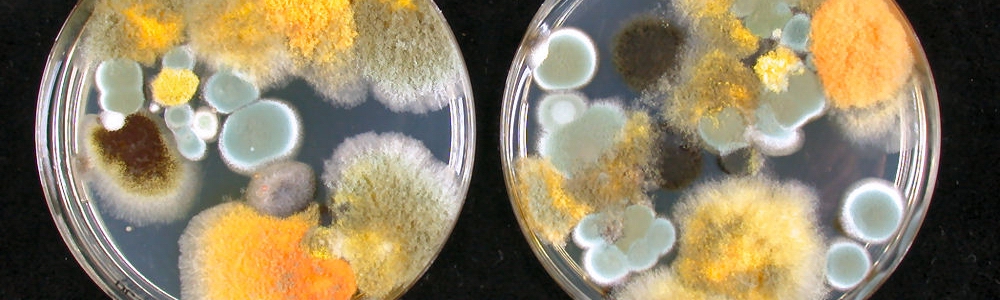
DNA extraction is a crucial step in any molecular assay. It can be difficult to obtain high quality and quantity genomic DNA from fungi, even from pure cultures. DNA extraction from complex matrixes, such as food, feed and other substrates, is even more difficult. The DNA extraction procedure can have a large effect on the outcome of the study and it is recommended to optimize this procedure for each matrix. One of the challenges is, similar to culture based techniques, to study a sufficient large sample because fungal growth can be heterogeneous through a sample (Geisen 2007). Another challenge is obtaining a sufficient quantity of DNA because low amounts can be present in the substrate and therefore a low copy number will be present in the final DNA extract. Besides the quantity, also the quality of the DNA should be high and compounds in the DNA extract can affect/inhibit the PCR amplification (Hayat et al. 2012). Various house-made DNA extraction methods are described and several commercial DNA extraction kits are available (Grube et al. 2015, Hohnadel et al. 2014, Rodriguez et al. 2012). It is evident that the DNA extraction procedure has an effect the microbial community composition that will detected. For example, DNA was extracted more efficiently after bead beating of 45 or 450 seconds, compared to no or a five seconds treatment (DeSantis et al. 2005). Fungi produce various microscopic structures that can react differently during DNA extraction and this directly challenges the efficiency of the procedure. DNA recovery will be different for each cell type. For example, food and indoor fungi can produce single celled structures (e.g. conidia in Penicillium, Aspergillus), multi-celled structures (e.g. Alternaria, Fusarium) and hyphae. These structures can be thin-walled and those cells are disrupted more easily than thick-walled and/or melanized cells, that are more difficult to disrupt, even by bead beating (Summerbell 2011).
PCR AND qPCR METHODS
PCR based methods are generally divided in three generations: conventional PCR (1st generation), real-time or quantitative PCR (qPCR; 2nd generation) and digital PCR (dPCR; 3rd generation). Conventional and real-time PCR based assays are most commonly applied to detect fungi in food and feed matrixes and can be rapid alternatives to traditional culturing and identification methods. The theoretic detection limit of PCR is down to 1 to 10 molecules, but this is not reached in practice. Factors influencing the detection glimit are e.g. the DNA extract quality, PCR conditions and primers. The presence of inhibitory substances or failure of a PCR can be checked using internal controls (Hoorfar et al. 2004). False negatives can be obtained when a primer or probe does not bind to variant sequences and false positives can be obtained when primers and probes bind to homologous sequences. Validation of the primers can be partially performed in silico using genome sequence data, but it remains a prerequisite challenging the assay with a large fungal diversity (Summerbell 2011). Digital PCR (dPCR; 3rd generation) can overcome difficulties encountered with qPCR, such as PCR inhibition and the use of an external standard curve, and is a promising tool when a low copy number needs to be detected (Rački et al. 2014). The application of dPCR in food and indoor mycology is limited and this paragraph will therefore not further deal with this technique.
In a conventional PCR assay, the result of the PCR is visualized at the end of the reaction, and in a qPCR assay the result is displayed after each amplification cycle. Conventional PCR therefore does not give any information on the initial copy number in the sample. The possibility to quantify the initial copy number using qPCR can be of added value when detection via conventional PCR is not sufficient. qPCR is in food mycology mainly applied for the detection of important mycotoxin-producing fungi (Atoui et al. 2007, Mayer et al. 2003, Nicolaisen et al. 2009), and the development of assays for spoilage fungi is more limited (Geisen 2007, Rawsthorne & Phister 2006). It should be stressed that quantification of food spoilage and indoor fungi using a qPCR assay has limitations. Targeting repetitive regions is unwanted because these occur in multiple copies per genome. For example, 40–240 ribosomal RNA genes (rDNA) copies per genome might occur (Griffin 1994, O’Donnell 1992). This bias can be overcome by targeting single copy genes (Geisen 2007). Another aspect is the variable number of nuclei in a fungal cell. Hyphae, conidiophores and even unicellular conidia can be uninucleate or multinucleate (Ishi et al. 2005, van Leeuwen et al. 2013) and this influences the correlation between the biomass present in a sample and qPCR data (Geisen 2007). PCR based assays are not designed for the detection of the general food mycobiota and only detect what they are designed for. A technical challenge is distinguishing dead and living material. A solution to detect only metabolically active and viable microorganisms is to target cDNA, which is generated via reverse transcription. This technique (RT-PCR) has been applied for the detection of viable fungi in wine (Hierro et al. 2006) and yogurts (Bleve et al. 2003).
NEXT GENERATION SEQUENCING
The use of next generation sequencing (NGS) in food sciences will be a great leap forward. NGS is a powerful tool and has the advantage of simultaneous identification of multiple contaminants, and it is even able to discover new and unexpected food spoilage organisms. The terms metagenomics and metagenetics are used in NGS. The term metagenomics refers to an approach, in which the total genomic content from a matrix is sequenced including taxonomic relevant genes as well as functional genes. In a metagenetics approach, the mycobiota of food and beverages is estimated by amplification of a marker. The most commonly used marker is one of the ITS regions (most often the ITS1 region); however, this region has limited power to resolve species, especially the species causing food spoilage and growing in indoor environments (e.g. Aspergillus, Cladosporium, Fusarium and Penicillium spp.). These species can share identical ITS sequences and other markers for species characterisation should be applied/investigated. In a metagenomics approach, other sequenced genes could be used for identification, resulting in information that is more accurate; however, this technique is more expensive and requires more knowledge on bioinformatics. Similarly as in PCR based techniques, the success of this methods also depends on the DNA quantity and quality. Disadvantages are the need of well-equipped and trained laboratories, and the impossibility to discriminate between metabolically live and dead micro-organisms (Cardinali et al. 2017, Rodriguez et al. 2015)